|







|
|
| |
The red mycelium pigmentation
of members of the Fusarium section Liseola is due to their production
of the monomeric naphthoquinone bikaverin (Leslie et al., 2004). Production
of this metabolite has in F. fujikuroi been linked to a 18 kb a gene
cluster (Figure 8). The cluster is regulated by a positive acting pathway
specific transcription factor (bik5) and positive acting NmrA-like regulator
(Wiemann et al., 2009).
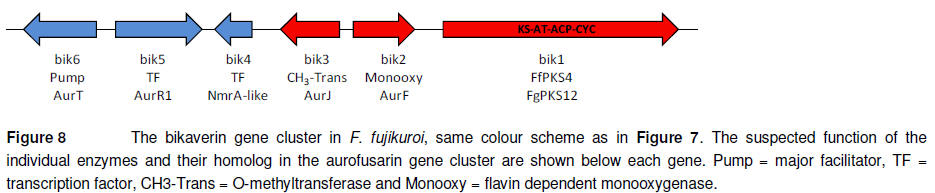
The cluster encodes three
catalytic enzymes: an O-methyltransferase (bik3), a flavin dependent
monooxygenase (bik2) and a non-reducing iPKS (bik1 or FfPKS4). Targeted
replacement of bik1 resulted in a down regulation of the gene cluster,
suggesting the existence of a feedback mechanism whereby the modifying
enzymes only are expressed if their substrates are present in the cell. A
theory that has gained further support as ectopic overexpression of bik1
(FfPKS4) in a Dbik1 strain restored cluster
expression, ruling out that the observed regulation was due to alterations
in the local chromatin structure in the Dbik1
strain (Wiemann et al., 2009). Similar down regulation effects have also
been reported for the cercosporin gene cluster in Cercospora nicotianae
(Chen et al., 2007). Down regulation of cluster expression was also
observed upon targeted replacement of bik2 and bik3, a situation that
unfortunately has hindered direct characterization of the intermediates in
the biosynthetic pathway that leads to bikaverin formation. However,
heterologous expression of bik1 (FfPKS4) in E. coli has shown that
the unmodified product of the iPKS is the compound SMA76a (Ma et al., 2007),
later confirmed by overexpression of the PKS in F. fujikuroi (Wiemann
et al., 2009).
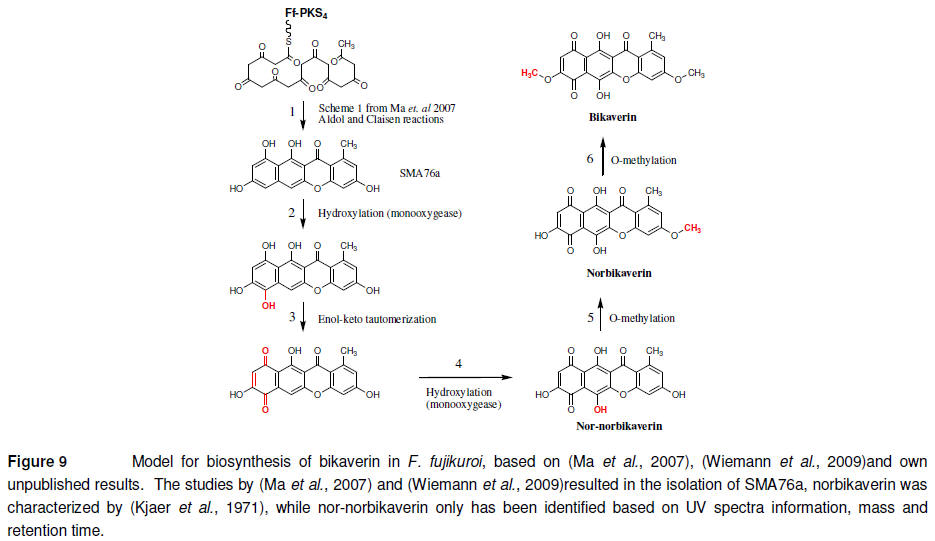
Though the bikaverin gene
cluster has been subjected to extensive analysis, a biosynthetic model for
the formation of the compound has not been presented. However, based on the
available data and theoretical biochemical considerations, I have formulated
one that fits with the reported chemical information as described in Figure
9. The order of step 2, 4, 5 and 6 are interchangeable and determination of
the correct order will require additional experimental data, generated by
heterologous expression of the entire biosynthetic pathway.
|
| |
|
References
 |
Leslie,J.F., Zeller,K.A., Logrieco,A., Mule,G.,
Moretti,A., and Ritieni,A. (2004) Species diversity of and toxin
production by Gibberella fujikuroi species complex strains isolated
from native Prairie Grasses in Kansas. Applied and Environmental
Microbiology 70: 2254-2262. |
 |
Wiemann,P., Willmann,A., Straeten,M., Kleigrewe,K.,
Beyer,M., Humpf,H.U., and Tudzynski,B. (2009) Biosynthesis of the red
pigment bikaverin in Fusarium fujikuroi: genes, their function and
regulation. Molecular Microbiology 72: 931-946. |
 |
Chen,H.Q., Lee,M.H., Daub,M.E., and Chung,K.R. (2007)
Molecular analysis of the cercosporin biosynthetic gene cluster in
Cercospora nicotianae. Molecular Microbiology 64: 755-770. |
 |
Ma,S.M., Zhan,J., Watanabe,K., Xie,X., Zhang,W.,
Wang,C.C., and Tang,Y. (2007) Enzymatic synthesis of aromatic polyketides
using PKS4 from Gibberella fujikuroi. Journal of the American
Chemical Society 129: 10642- |
 |
Kjaer,D., Kjaer,A., Pedersen,C., Bulock,J.D., and
Smith,J.R. (1971) Bikaverin and Norbikaverin, Benzoxanthentrione Pigments
of Gibberella-Fujikuroi. Journal of the Chemical Society C-Organic:
2792-. |
|
