Sexual crosses
The following protocol is intented for providing a
gentic confromation that a observed phenotype of a constructed deletion
strain (using hygromycin resistance) is caused by the introduced mutation (essentially
checking for unintented and unmarked secondary mutations in the strain). The
cross of the mutant and the tester (here wild type) should show that the
observed phenotype is 100% linked to the hygromycin resistance.
However, with small modifications the protocol can also
be used for construction of double deletion strains by varying the selection pressure and assay conditions.
Protocol
-
Place mycelial plugs (3-4 mm) of the two strains you
want to cross on a thick 9
cm carrot agar plate. One strain should be HygromycinS, the other HygromycinR. Make 5
plates for each cross.

-
Incubate at 20 oC under (ca. 25 cm) a
mixture of continuous cool white and black-light fluorescent lighting. To
secure high humidity the plates are incubated in a clear plastic box with
a lid, and a beaker of water is placed inside.
-
After approx. 7 days of growth, when mycelia meet at the
middle, apply 2 ml of 2.5% aqueous Tween 60 solution and spread with a
Drigalsky spatula to flatten the mycelium.

Incubate
for an additional 10 – 14 days.
-
Now perithecia has formed.

-
-
-
-
-
-
-
Pick a perithecium along the line between the two
strains with a pair of
forceps.
Transfer the perithecium to an
Eppendorf tube with 400 µl sterile MilliQ water and squeeze the perithecium
using the pair of forceps to release the ascospores from the perithecium.
Vortex briefly and transfer 20 µl
spore suspension to a microscope slide to check the ascospore quality in a
microscope.
-
Add 600 µl sterile MilliQ water to the spore suspension
and filter through a sterile
pasteur pipette with a bit of glass wool. The
glass wool will hold back any hyphae but let ascospores pass.
-
Divide the spore suspension outcome in two Eppendorf
tubes. Plate 20 µl portions from one tube on 14 cm PDA agar plates and
from the other tube on 9 cm PDA hyg75 agar plates. Spread with
a Drigalsky spatula.
Incubate at 20 oC for 3
days to allow spore germination.
-
Count colonies on the PDA+/- hyg to determine if it is
a cross.
-
Draw a grid of 6 x 6 lines on 9 cm PDA+/- hyg150
agar plates.
Transfer all colonies from the 14 cm
PDA agar plates to PDA and PDA hyg150 agar plates using s
sterile toothpick. There is room for 34 colonies on each plate.
Incubate at 20 oC for 4
days.
-
Score progeny for Hygromycin resistance and colour
variation.
PDA
PDAhyg75
After 2 days 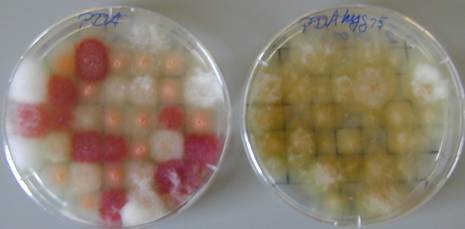
PDA
PDAhyg75
After 5 days
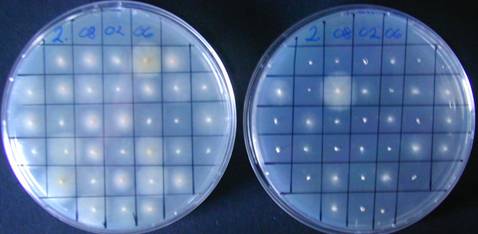
Preparing the carrot-agar (Klittitch 1988)
-
Chop 400 g of baby carrots in a
food processor
-
Place the carrots in a 2 L
glass beaker, add 400 ml deionised water, cover loosely with Saran wrap
and microwave 2 times at 10 minutes
-
Let it cool for 1 to 2 hrs
-
Puree in a food processor
-
Return to beaker and add 500 ml
water, mix well
-
Dispense evenly into 5 x 500 ml
Pyrex bottles (ap. 225 ml per bottle) containing 4 g agar per bottle (ap.
1.8 % agar)
-
Autoclave for 20 minutes.
Tween 60 solution, 2.5 %
Tween 60 stock is highly viscous;
microwave about 20 seconds to liquefy. Using wide-bore pippet, transfer 3.75
ml to a 250 ml Pyrex bottle containing 146.5 ml warmed deionised water.
Autoclave.

